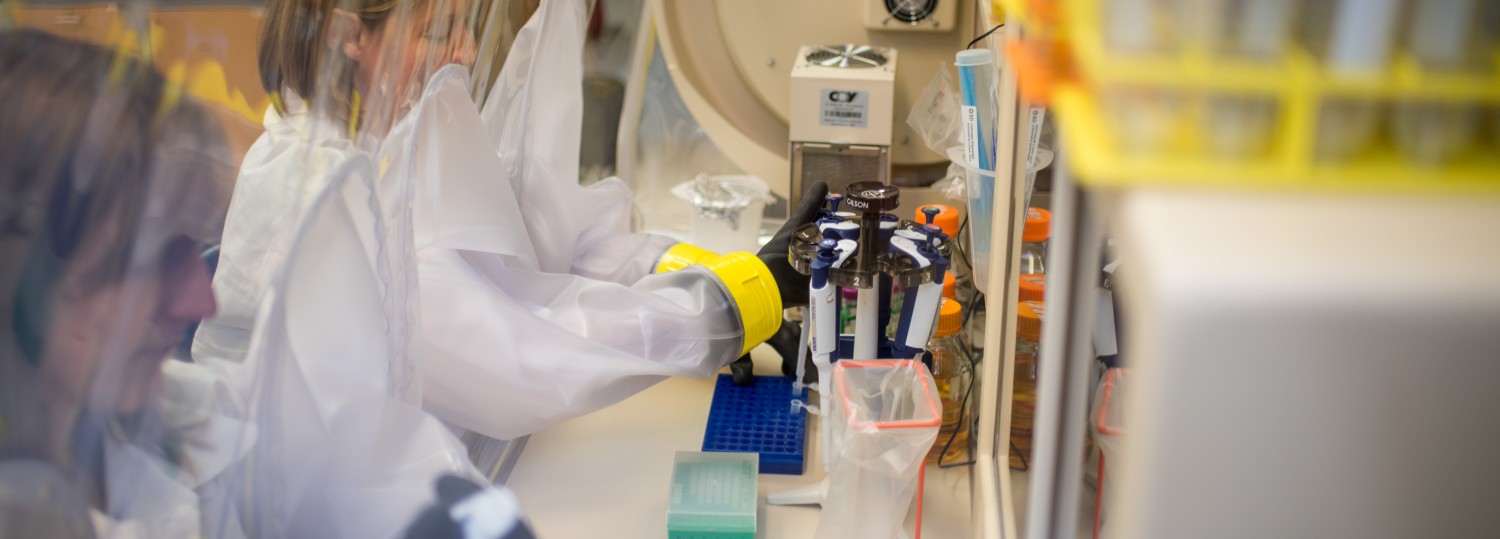
Research
Our research focuses on defining the gastrointestinal tract microbiome and metabolome during different disease states to understand how perturbations to this system affect animal and human health. Colonization resistance describes the ability of the gastrointestinal tract to prevent colonization by pathogens. Currently, we are exploring the interactions between the gut microbiota and the pathogen Clostridium difficile, a significant and re-emerging public health problem. C. difficile infection (CDI) is the leading nosocomial infection in the United States and is also becoming an emerging threat worldwide.
Our long-term research interests include studying infectious diseases and how they impact human health. A major goal of our work is to create an integrated model of the complex interactions among the gut microbiota, pathogen, and host. To accomplish this, we are integrating data obtained from high-throughput methods that analyze the gut microbiome, the metabolome and host immune responses in animal models and humans. It will be critically important to employ a One Health systems biology approach to defining alterations in the gastrointestinal tract following antibiotic exposure.
With the advent of “omics” technologies, complex communities such as the gastrointestinal tract can be defined. We can also explore how the metabolism of nutrients, by members of the indigenous gut microbiota, alters colonization resistance to C. difficile and other pathogens. Alterations in the gut microbiota have been associated with many disease states including diabetes, obesity, irritable bowel disease, and other metabolic disorders. Defining the role that bacteria play in the gut will allow us to design targeted approaches for prevention and treatment of these gastrointestinal disorders.
